pacman::p_load(ggstatsplot, tidyverse)Hands-on_Ex04
Visual Statistical analysis
Getting Started
Install and launch R packages
Import data
exam <- read_csv("data/Exam_data.csv")One-sample test: gghistostats() method
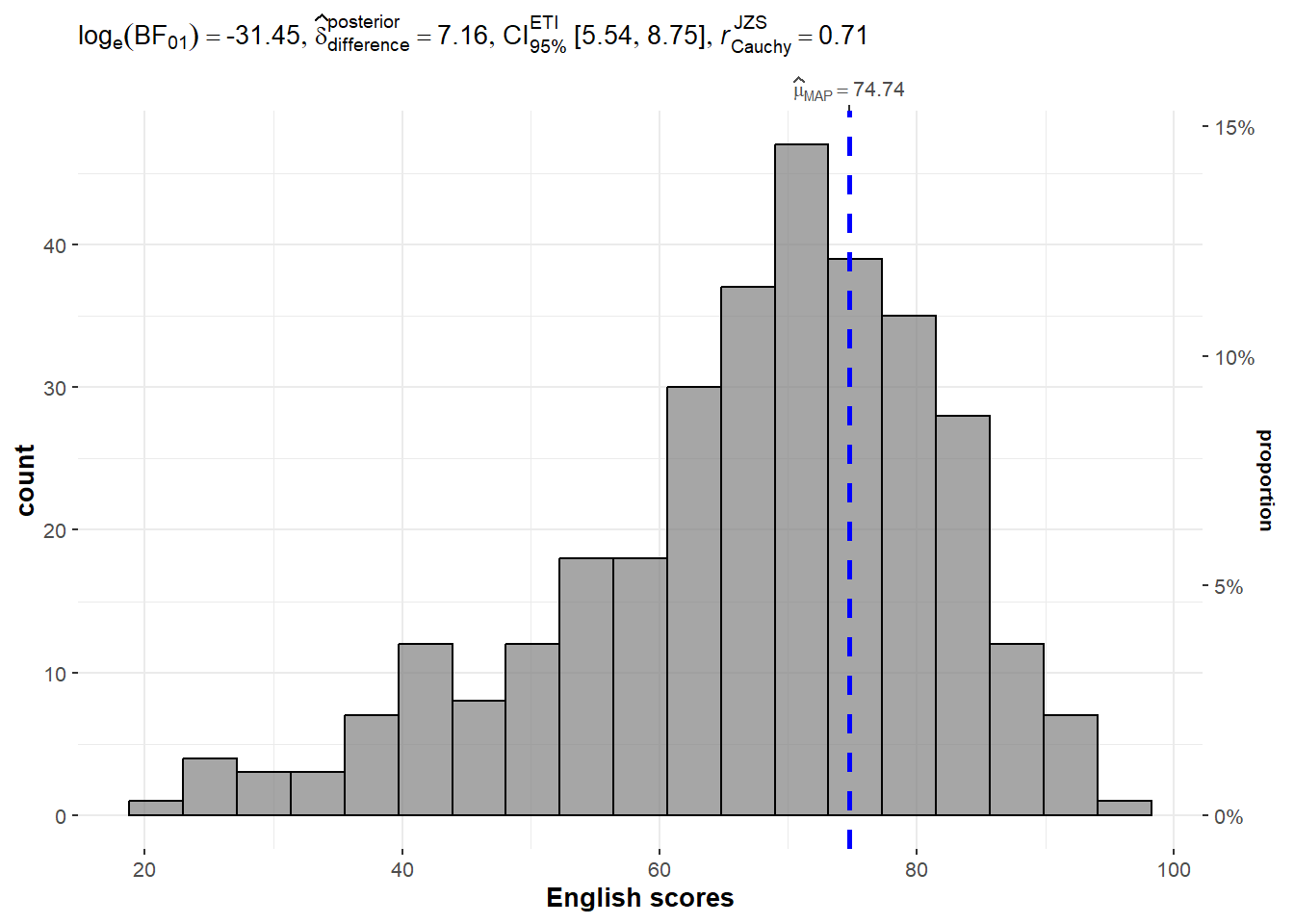
A Bayes factor is the ratio of the likelihood of one particular hypothesis to the likelihood of another. It can be interpreted as a measure of the strength of evidence in favor of one theory among two competing theories.
set.seed(1234)
gghistostats(
data = exam,
x = ENGLISH,
type = "bayes",
test.value = 60,
xlab = "English scores"
)
Two-sample mean test: ggbetweenstats() method
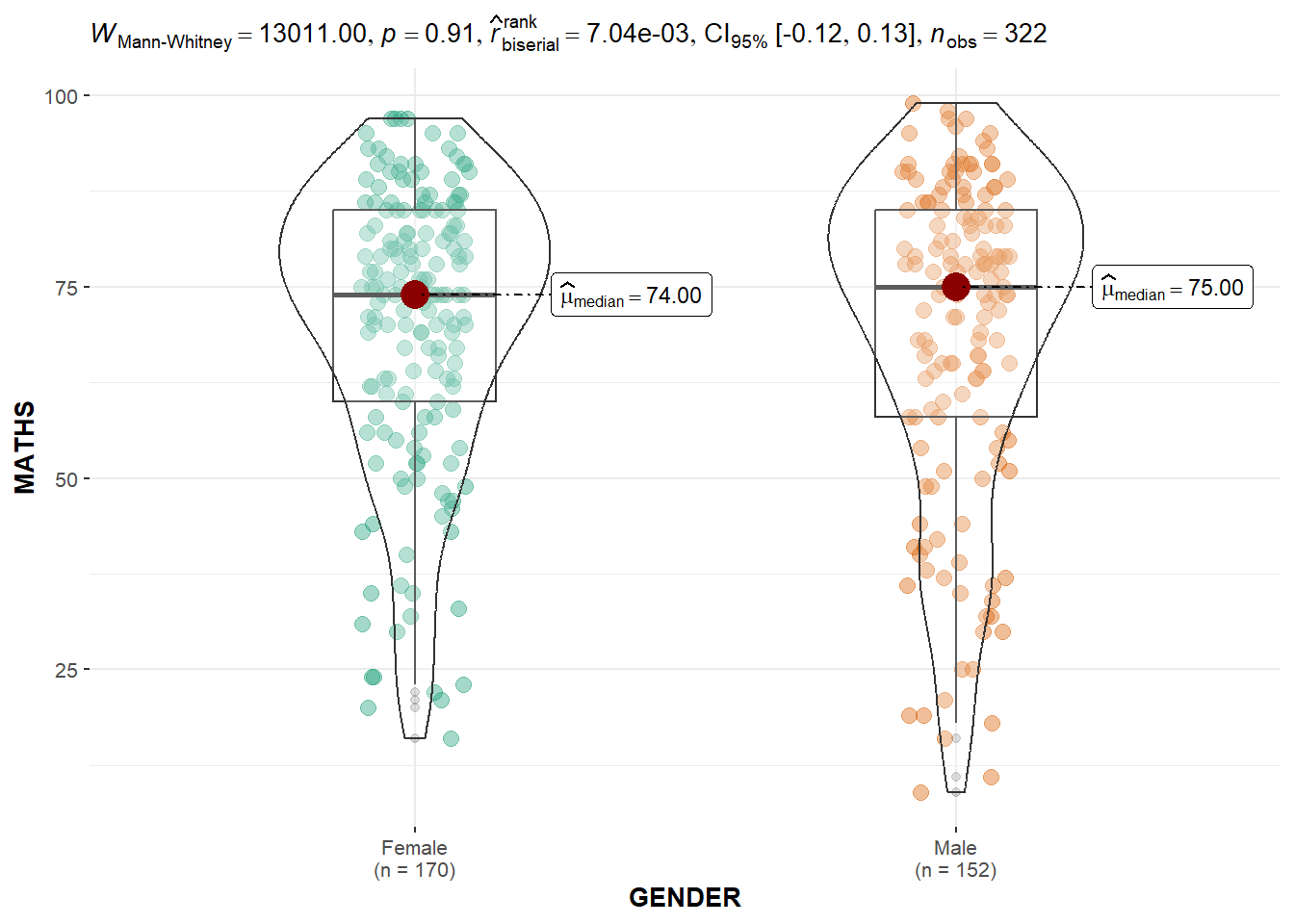
Two-sample mean test of Maths scores by gender.
ggbetweenstats(
data = exam,
x = GENDER,
y = MATHS,
type = "np",
messages = FALSE
)
“ns” → only non-significant “s” → only significant “all” → everything
Oneway ANOVA Test: ggbetweenstats() method
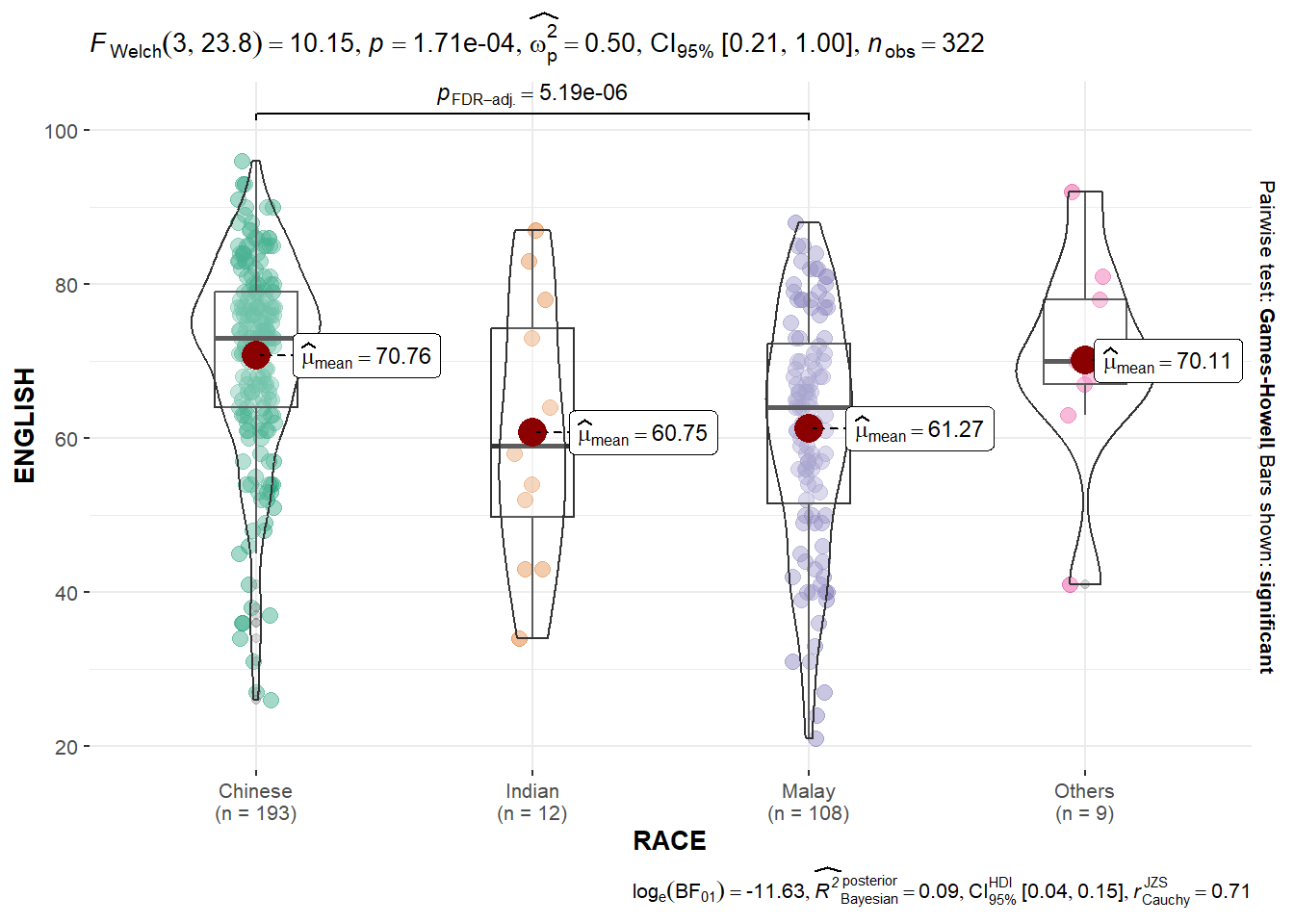
One-way ANOVA test on English score by race.
ggbetweenstats(
data = exam,
x = RACE,
y = ENGLISH,
type = "p",
mean.ci = TRUE,
pairwise.comparisons = TRUE,
pairwise.display = "s",
p.adjust.method = "fdr",
messages = FALSE
)
Significant Test of Correlation: ggscatterstats()
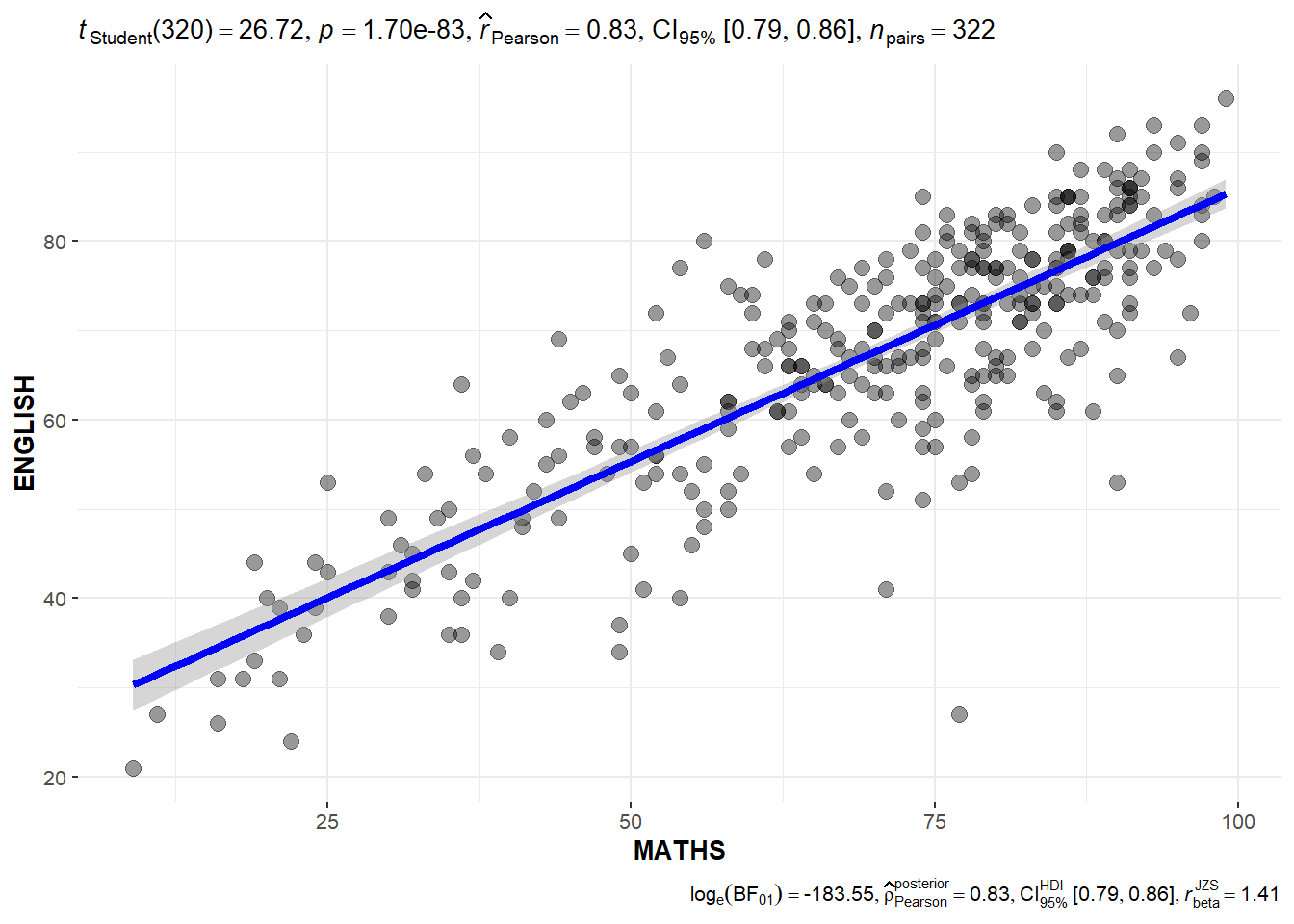
Significant Test of Correlation between Maths scores and English scores.
ggscatterstats(
data = exam,
x = MATHS,
y = ENGLISH,
marginal = FALSE,
)
Significant Test of Association: ggbarstats() methods
Maths scores is binned into a 4-class variable by using cut().
exam1 <- exam %>%
mutate(MATHS_bins =
cut(MATHS,
breaks = c(0,60,75,85,100))
)Significant Test of Association between Maths score and gender.
ggbarstats(exam1,
x = MATHS_bins,
y = GENDER)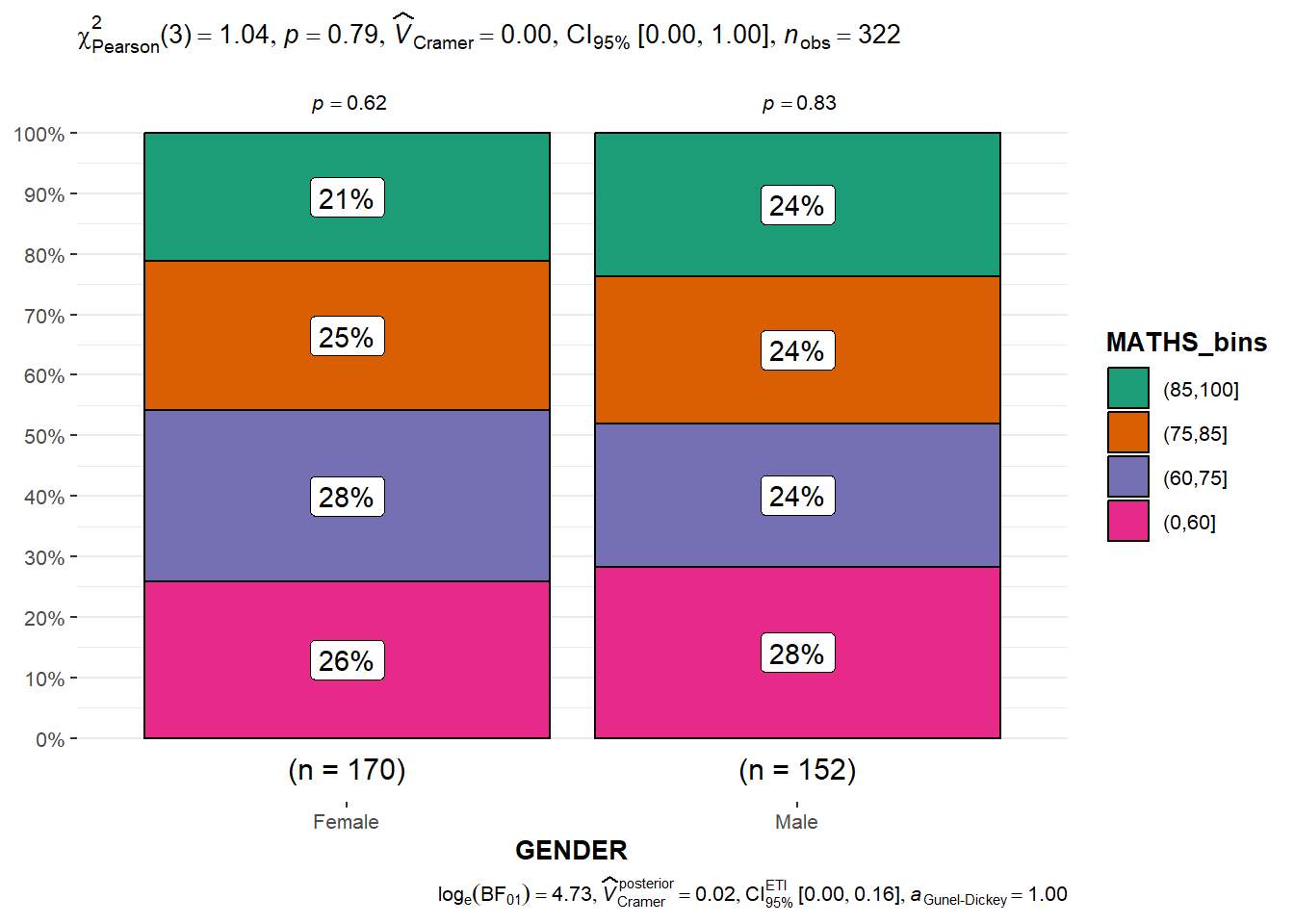
Visualizing Models
Toyota Corolla case study will be used. The purpose of study is to build a model to discover factors affecting prices of used-cars by taking into consideration a set of explanatory variables.
Install and load neccessary libraries and import dataset
pacman::p_load(readxl, performance, parameters, see)
car_resale <- read_xls("data/ToyotaCorolla.xls",
"data")
car_resale# A tibble: 1,436 × 38
Id Model Price Age_08_04 Mfg_Month Mfg_Year KM Quarterly_Tax Weight
<dbl> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 81 TOYOTA … 18950 25 8 2002 20019 100 1180
2 1 TOYOTA … 13500 23 10 2002 46986 210 1165
3 2 TOYOTA … 13750 23 10 2002 72937 210 1165
4 3 TOYOTA… 13950 24 9 2002 41711 210 1165
5 4 TOYOTA … 14950 26 7 2002 48000 210 1165
6 5 TOYOTA … 13750 30 3 2002 38500 210 1170
7 6 TOYOTA … 12950 32 1 2002 61000 210 1170
8 7 TOYOTA… 16900 27 6 2002 94612 210 1245
9 8 TOYOTA … 18600 30 3 2002 75889 210 1245
10 44 TOYOTA … 16950 27 6 2002 110404 234 1255
# ℹ 1,426 more rows
# ℹ 29 more variables: Guarantee_Period <dbl>, HP_Bin <chr>, CC_bin <chr>,
# Doors <dbl>, Gears <dbl>, Cylinders <dbl>, Fuel_Type <chr>, Color <chr>,
# Met_Color <dbl>, Automatic <dbl>, Mfr_Guarantee <dbl>,
# BOVAG_Guarantee <dbl>, ABS <dbl>, Airbag_1 <dbl>, Airbag_2 <dbl>,
# Airco <dbl>, Automatic_airco <dbl>, Boardcomputer <dbl>, CD_Player <dbl>,
# Central_Lock <dbl>, Powered_Windows <dbl>, Power_Steering <dbl>, …Multiple Regression Model using lm()
model <- lm(Price ~ Age_08_04 + Mfg_Year + KM +
Weight + Guarantee_Period, data = car_resale)
model
Call:
lm(formula = Price ~ Age_08_04 + Mfg_Year + KM + Weight + Guarantee_Period,
data = car_resale)
Coefficients:
(Intercept) Age_08_04 Mfg_Year KM
-2.637e+06 -1.409e+01 1.315e+03 -2.323e-02
Weight Guarantee_Period
1.903e+01 2.770e+01 Model Diagnostic: checking for multicolinearity:
Use check_collinearity() of performance package
check_collinearity(model)# Check for Multicollinearity
Low Correlation
Term VIF VIF 95% CI Increased SE Tolerance Tolerance 95% CI
KM 1.46 [ 1.37, 1.57] 1.21 0.68 [0.64, 0.73]
Weight 1.41 [ 1.32, 1.51] 1.19 0.71 [0.66, 0.76]
Guarantee_Period 1.04 [ 1.01, 1.17] 1.02 0.97 [0.86, 0.99]
High Correlation
Term VIF VIF 95% CI Increased SE Tolerance Tolerance 95% CI
Age_08_04 31.07 [28.08, 34.38] 5.57 0.03 [0.03, 0.04]
Mfg_Year 31.16 [28.16, 34.48] 5.58 0.03 [0.03, 0.04]check_c <- check_collinearity(model)
plot(check_c)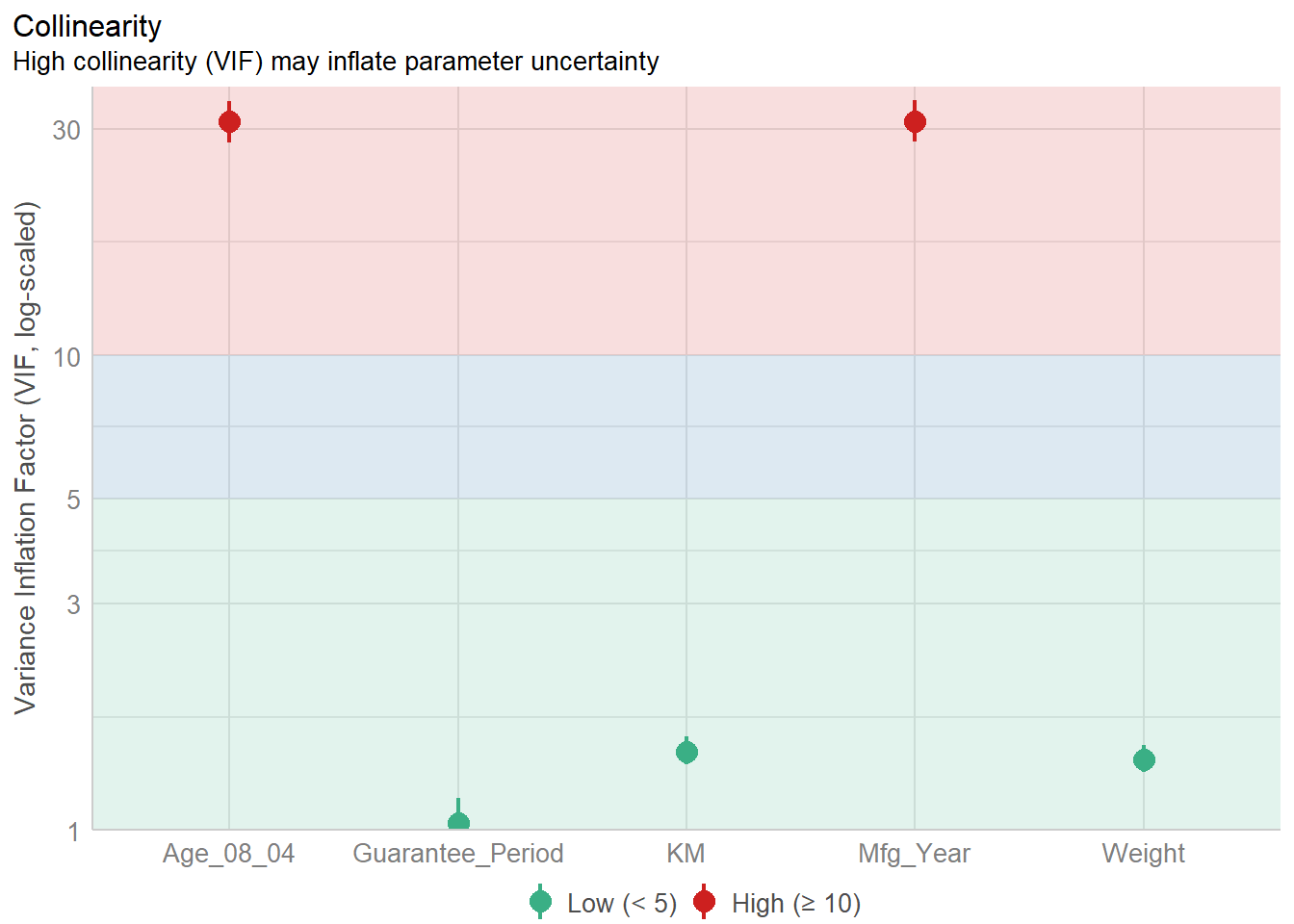
Model Diagnostic: checking normality assumption
Use check_normality() of performance package
model1 <- lm(Price ~ Age_08_04 + KM +
Weight + Guarantee_Period, data = car_resale)
check_n <- check_normality(model1)
plot(check_n)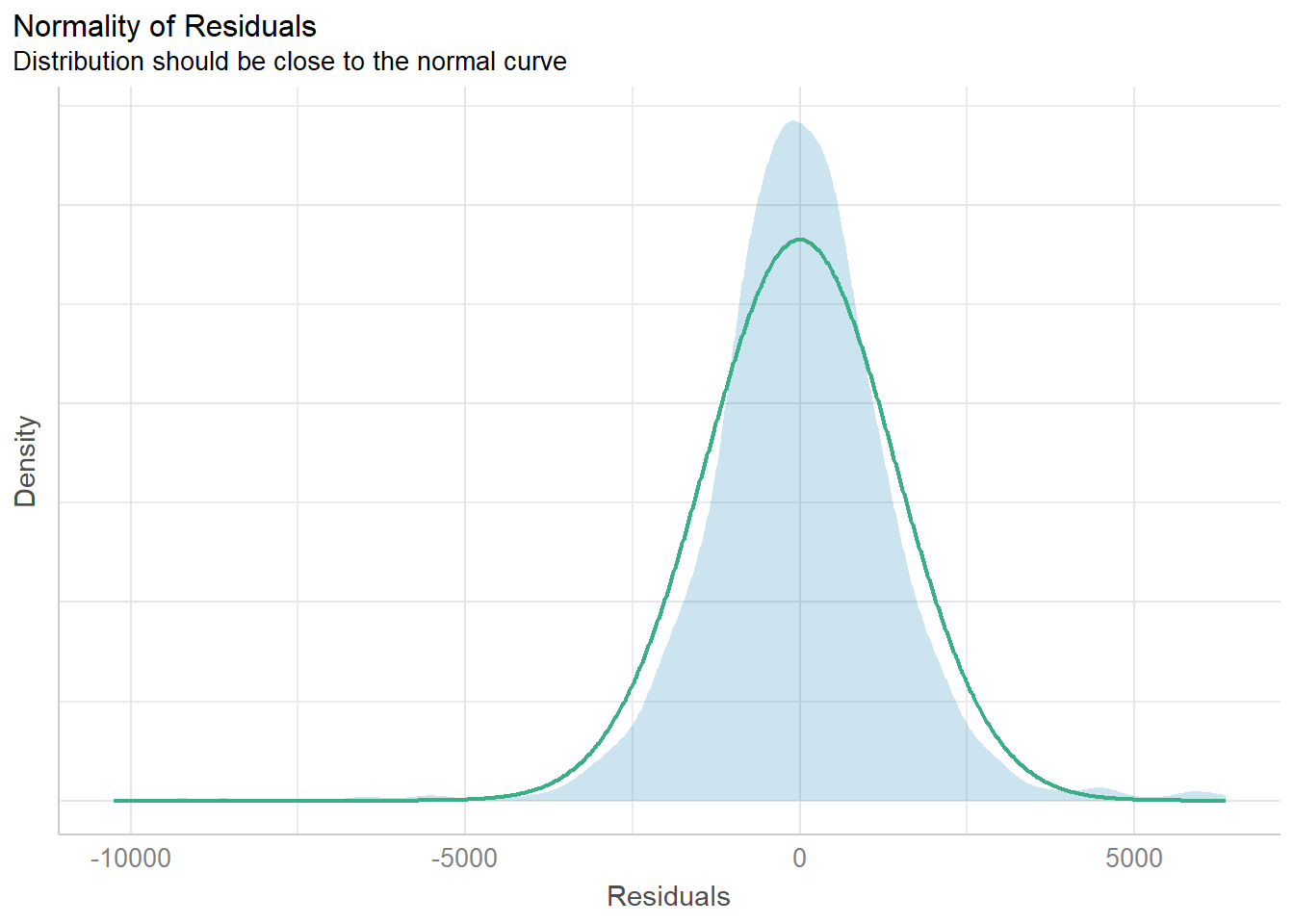
Model diagnostic: Check model for homogeneity of variances
Use check_heteroscedasticity() of performance package
check_h <- check_heteroscedasticity(model1)
plot(check_h)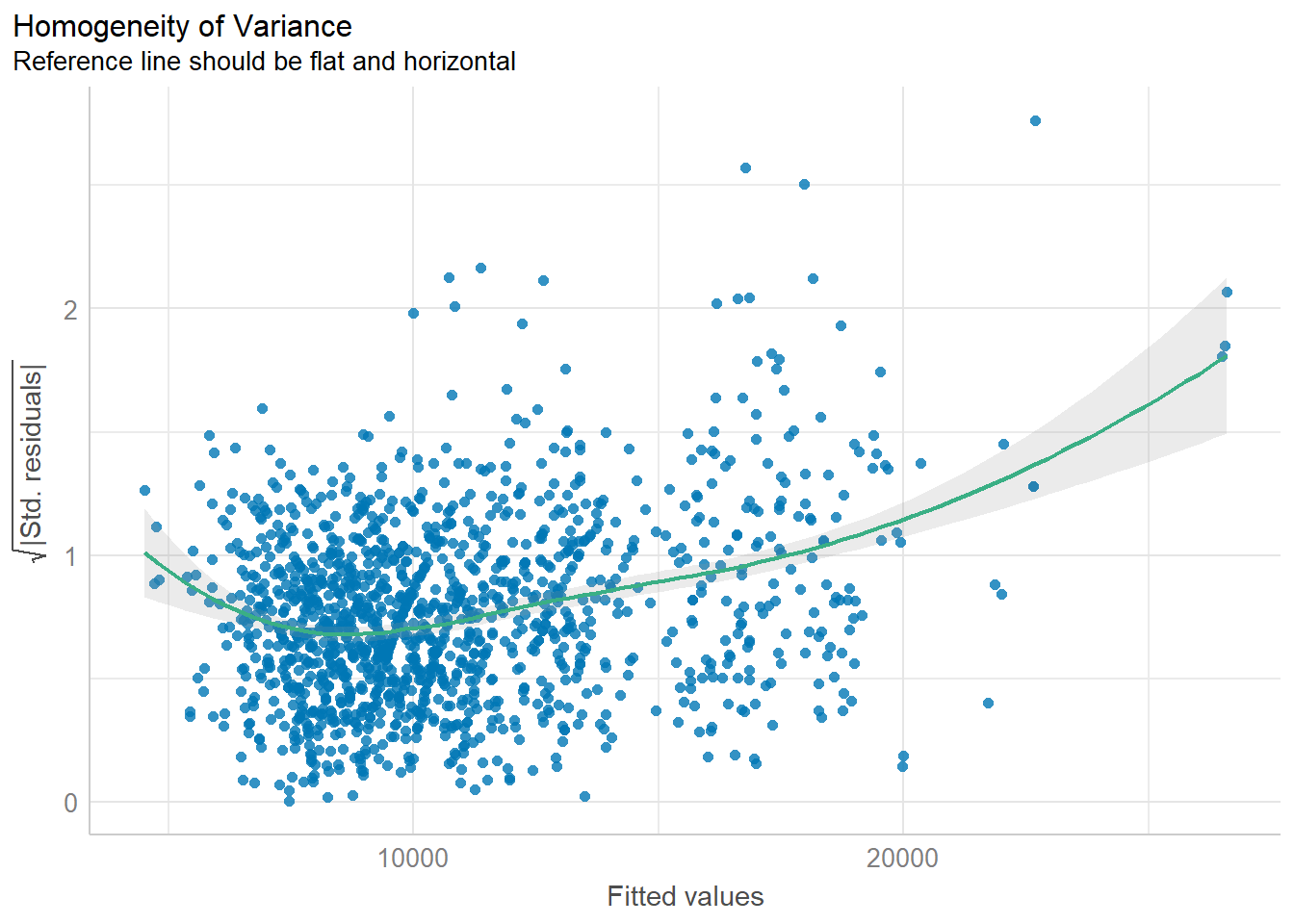
Model Diagnostic: Complete check
Use check_model().
check_model(model1)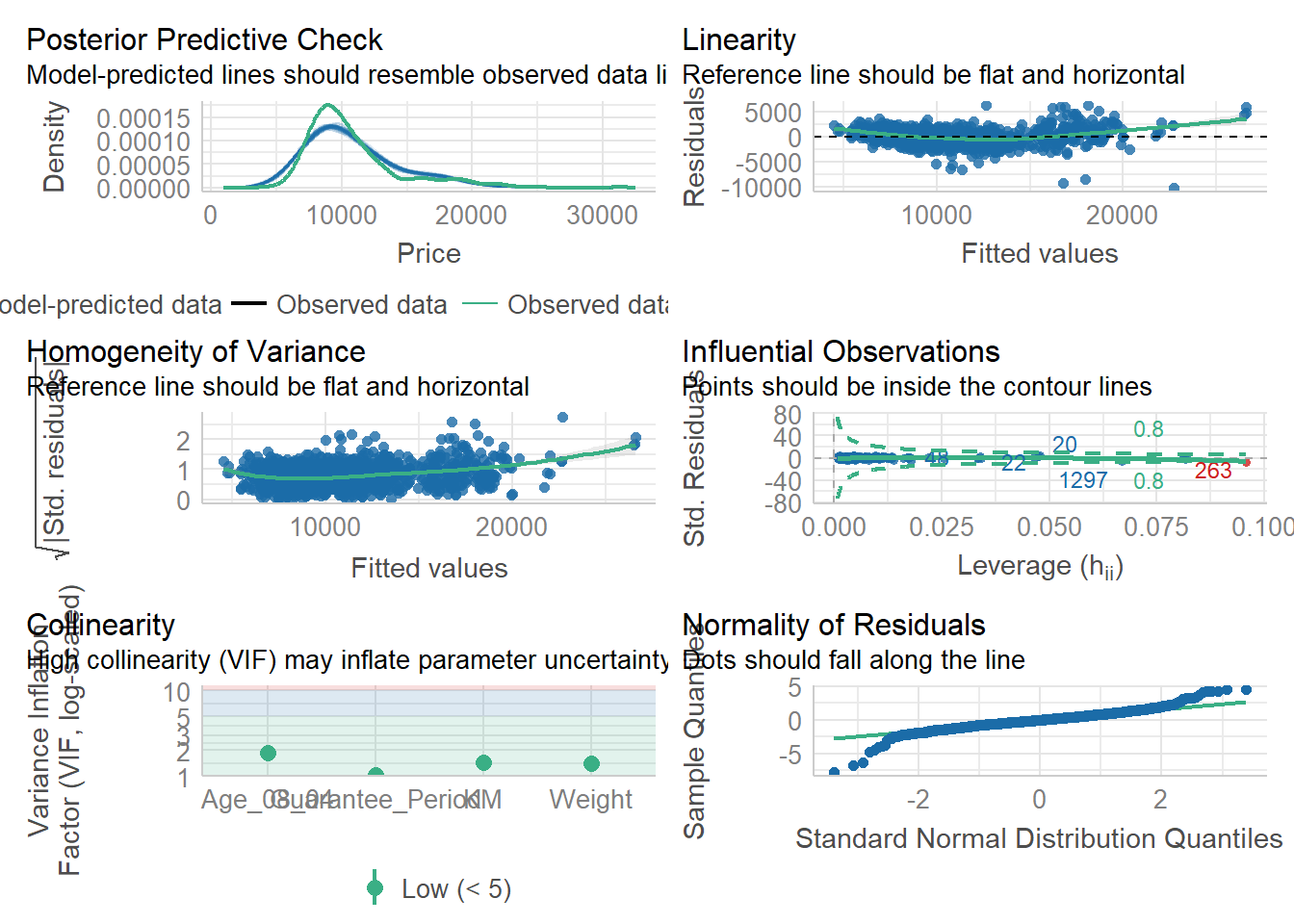
Visualising Regression Parameters: see methods
plot() of see package and parameters() of parameters package is used to visualise the parameters of a regression model.
plot(parameters(model1))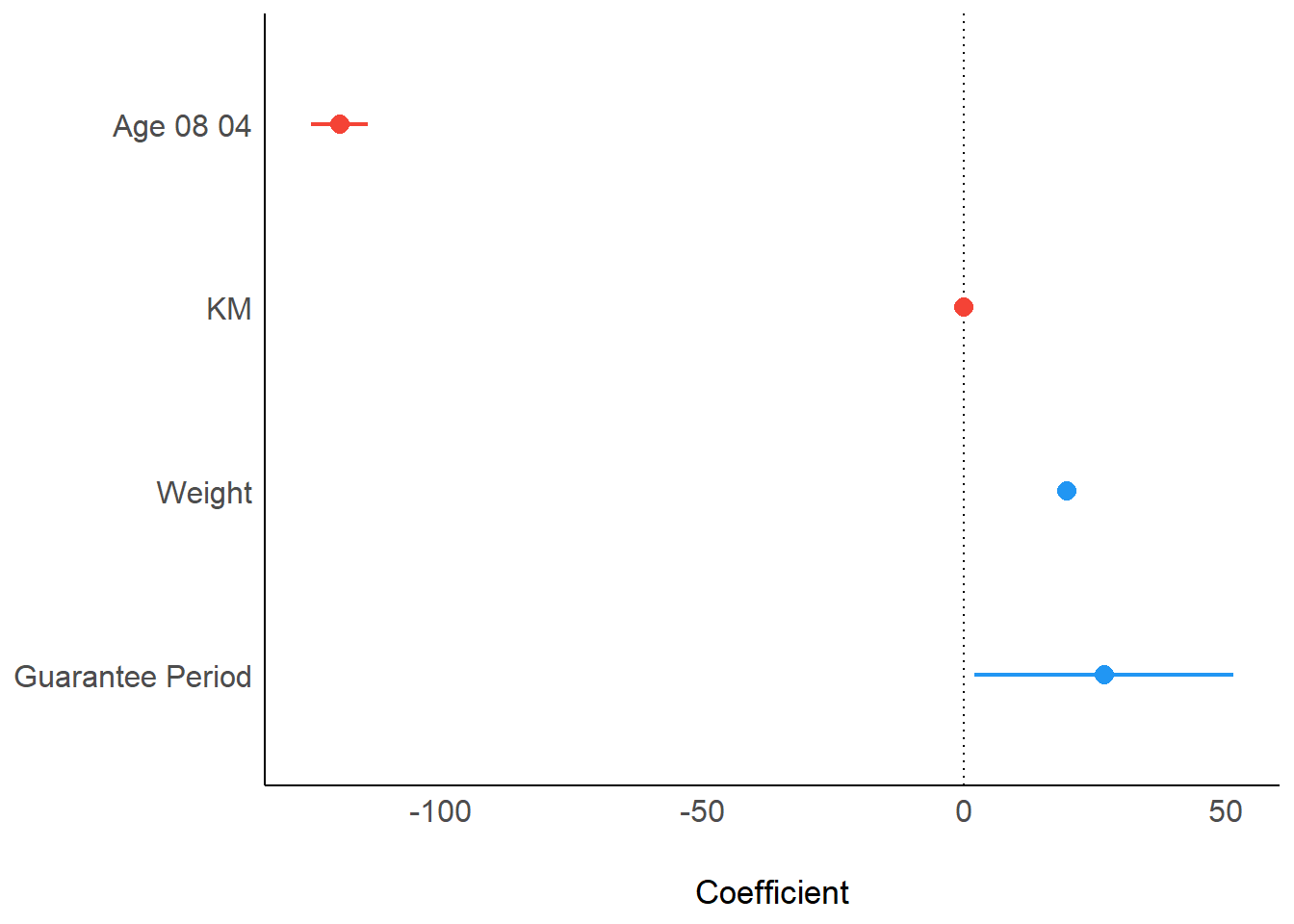
Visualising Regression ParametersL ggcoefstats() methods
Use ggcoefstats() of ggstatsplot package to visualise the parameters of a regression model
ggcoefstats(model1,
output = "plot")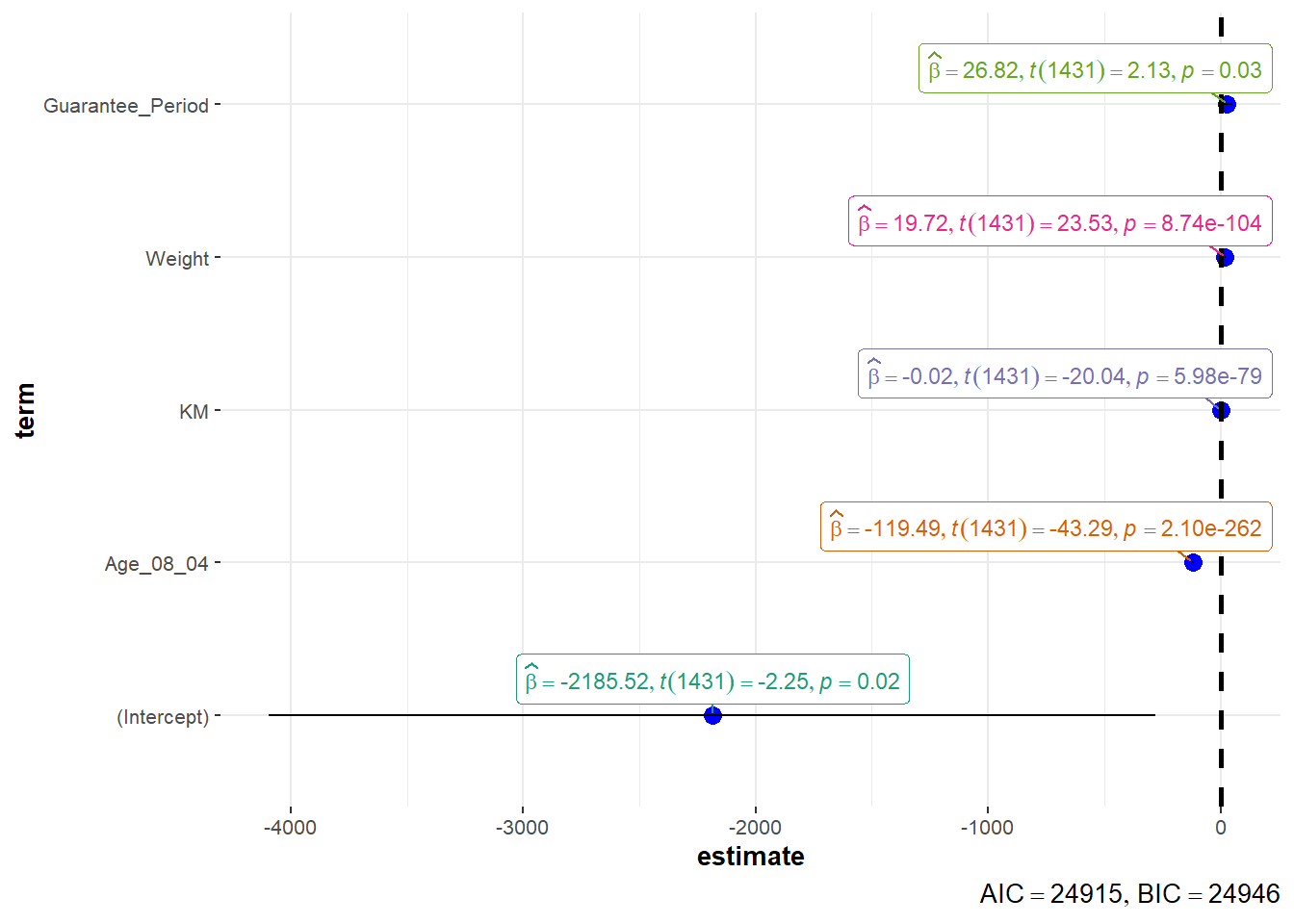
Visualizing Uncertainty
Visualizing Uncertainty of Point Estimates
pacman::p_load(tidyverse, plotly, crosstalk, DT, ggdist, gganimate)
exam <- read_csv("data/Exam_data.csv")Visualizing uncertainty of point estimates: ggplot2 methods
my_sum <- exam %>%
group_by(RACE) %>%
summarise(
n=n(),
mean=mean(MATHS),
sd=sd(MATHS)
) %>%
mutate(se=sd/sqrt(n-1))
knitr::kable(head(my_sum), format = 'html')| RACE | n | mean | sd | se |
|---|---|---|---|---|
| Chinese | 193 | 76.50777 | 15.69040 | 1.132357 |
| Indian | 12 | 60.66667 | 23.35237 | 7.041005 |
| Malay | 108 | 57.44444 | 21.13478 | 2.043177 |
| Others | 9 | 69.66667 | 10.72381 | 3.791438 |
Plot the standard error of mean maths score by race.
ggplot(my_sum) +
geom_errorbar(
aes(x=reorder(RACE,-mean),
ymin=mean-se,
ymax=mean+se),
width=0.2,
colour="black",
alpha=0.9,
size=0.5) +
geom_point(aes
(x=RACE,
y=mean),
stat="identity",
color="red",
size = 1.5,
alpha=1) +
labs(x="Race") +
ggtitle("Standard error of mean
maths score by race")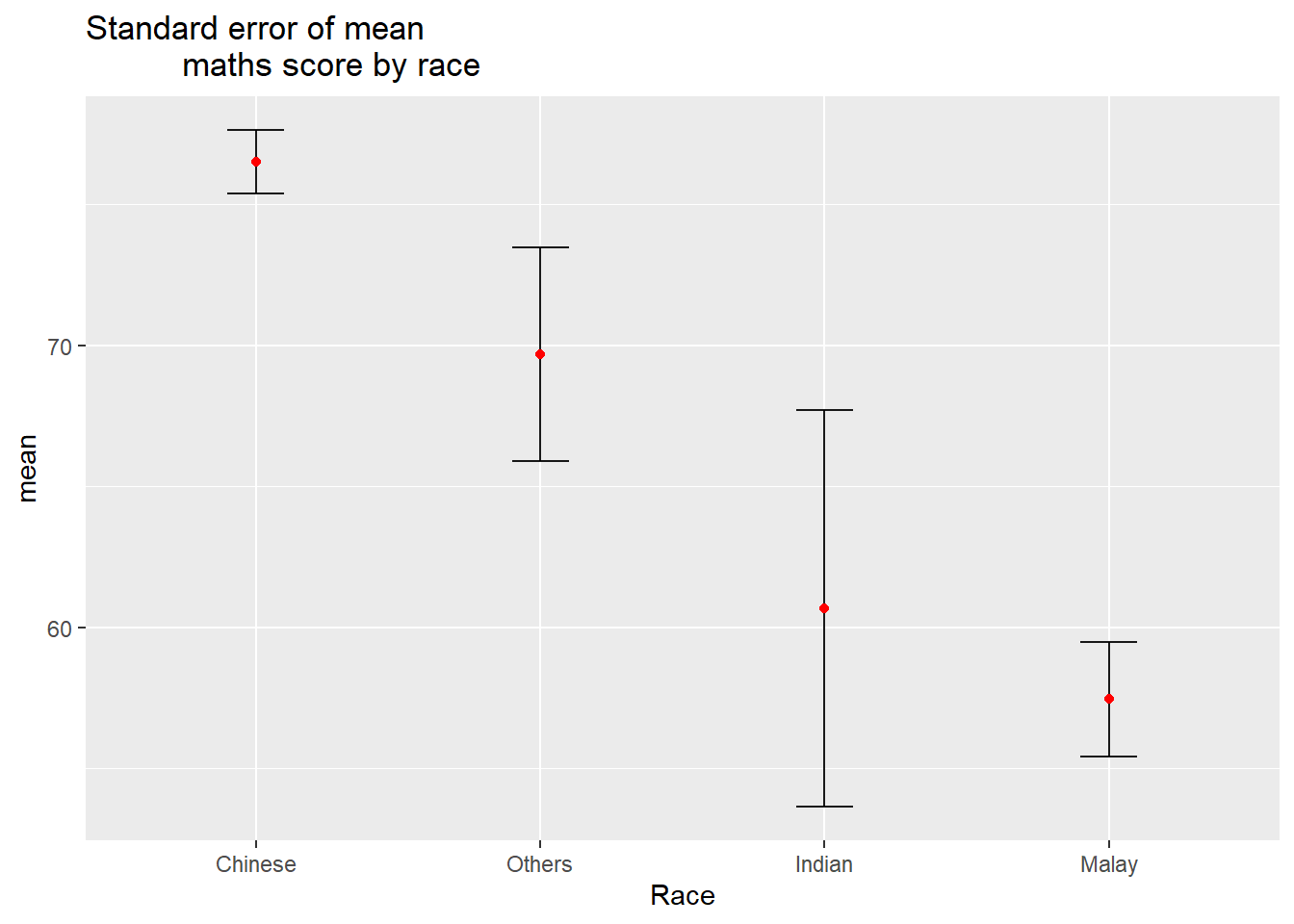
Plot the 95% confidence interval of mean maths score by race.
ggplot(my_sum) +
geom_errorbar(
aes(x=reorder(RACE,-mean),
ymin=mean-1.96*se,
ymax=mean+1.96*se),
width=0.2,
colour="black",
alpha=0.9,
size=0.5) +
geom_point(aes
(x=RACE,
y=mean),
stat="identity",
color="red",
size = 1.5,
alpha=1) +
labs(x="Race") +
ggtitle("95% confidence interval of mean
maths score by race")+
theme_minimal()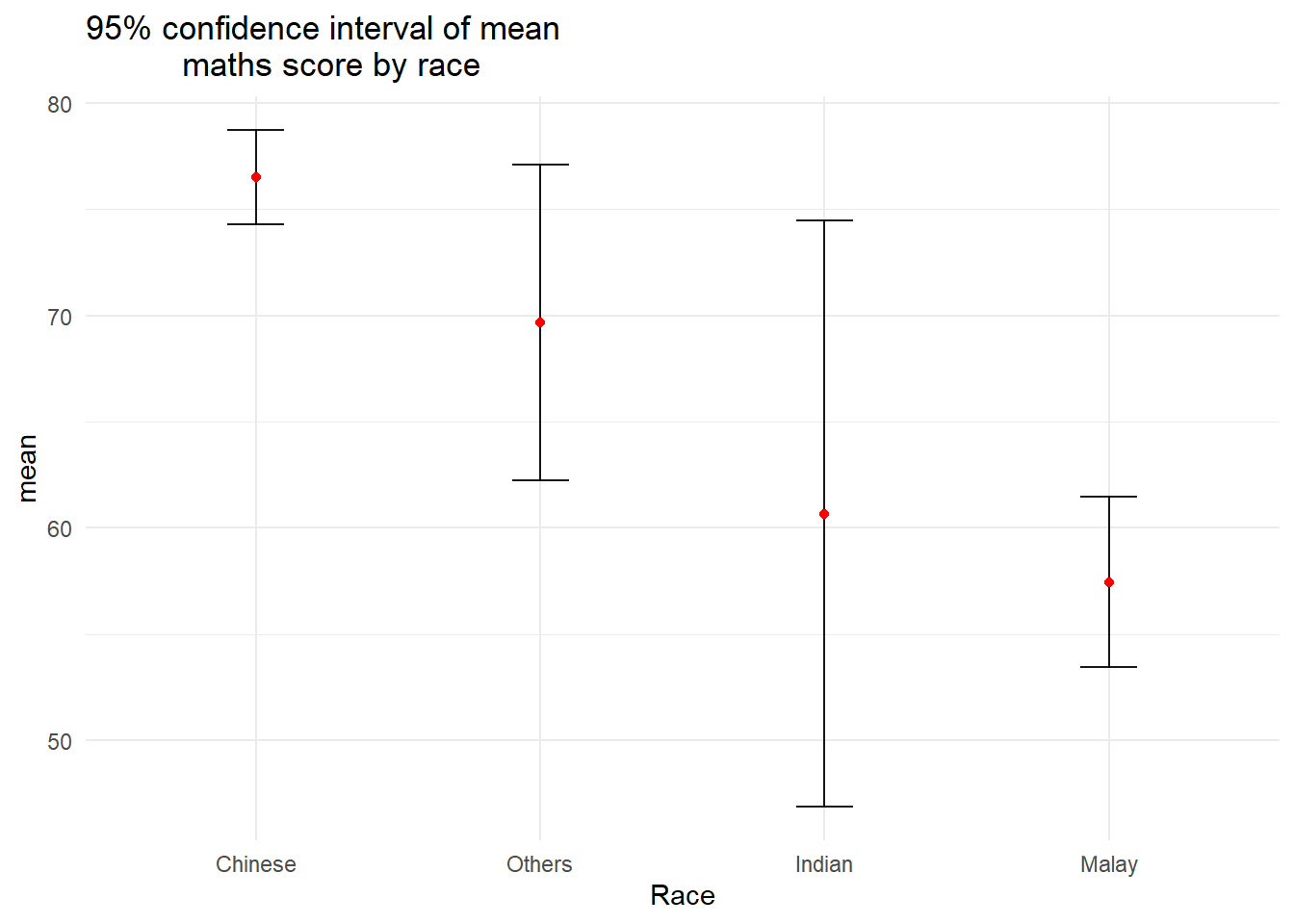
Plot interactive error bars for the 99% confidence interval of mean maths score by race.
d <- highlight_key(my_sum)
p <- ggplot(d) +
geom_errorbar(
aes(x=reorder(RACE,-mean),
ymin=mean-2.58*se,
ymax=mean+2.58*se),
width=0.2,
colour="black",
alpha=0.9,
size=0.5) +
geom_point(aes
(x=RACE,
y=mean,
text=paste("Race:", RACE,
"<br>N:", n,
"<br>Avg. Scores:", round(mean, digits = 2),
"<br>99% CI:[", round(mean - 2.58*se, digits = 2), ", ", round(mean + 2.58*se, digits = 2), "]")),
stat="identity",
color="red",
size = 1.5,
alpha=1) +
labs(x="Race") +
ggtitle("99% confidence interval of mean
maths score by race")+
theme_minimal()
gg <- highlight(ggplotly(p, tooltip="text"),
"plotly_selected")
dt <- DT::datatable(d,
colnames = c("","No. of pupils", "Avg Scores", "Std Dev", "Std Error")) |>
formatRound(columns = c("mean", "sd", "se"), digits = 2)
crosstalk::bscols(gg,
dt,
widths = 5) Visualising Uncertainty: ggdist package
stat_pointinterval() of ggdist is used to build a visual for displaying distribution of maths scores by race.
exam %>%
ggplot(aes(x = RACE,
y = MATHS)) +
stat_pointinterval() + #<<
labs(
title = "Visualising confidence intervals of mean math score",
subtitle = "Mean Point + Multiple-interval plot")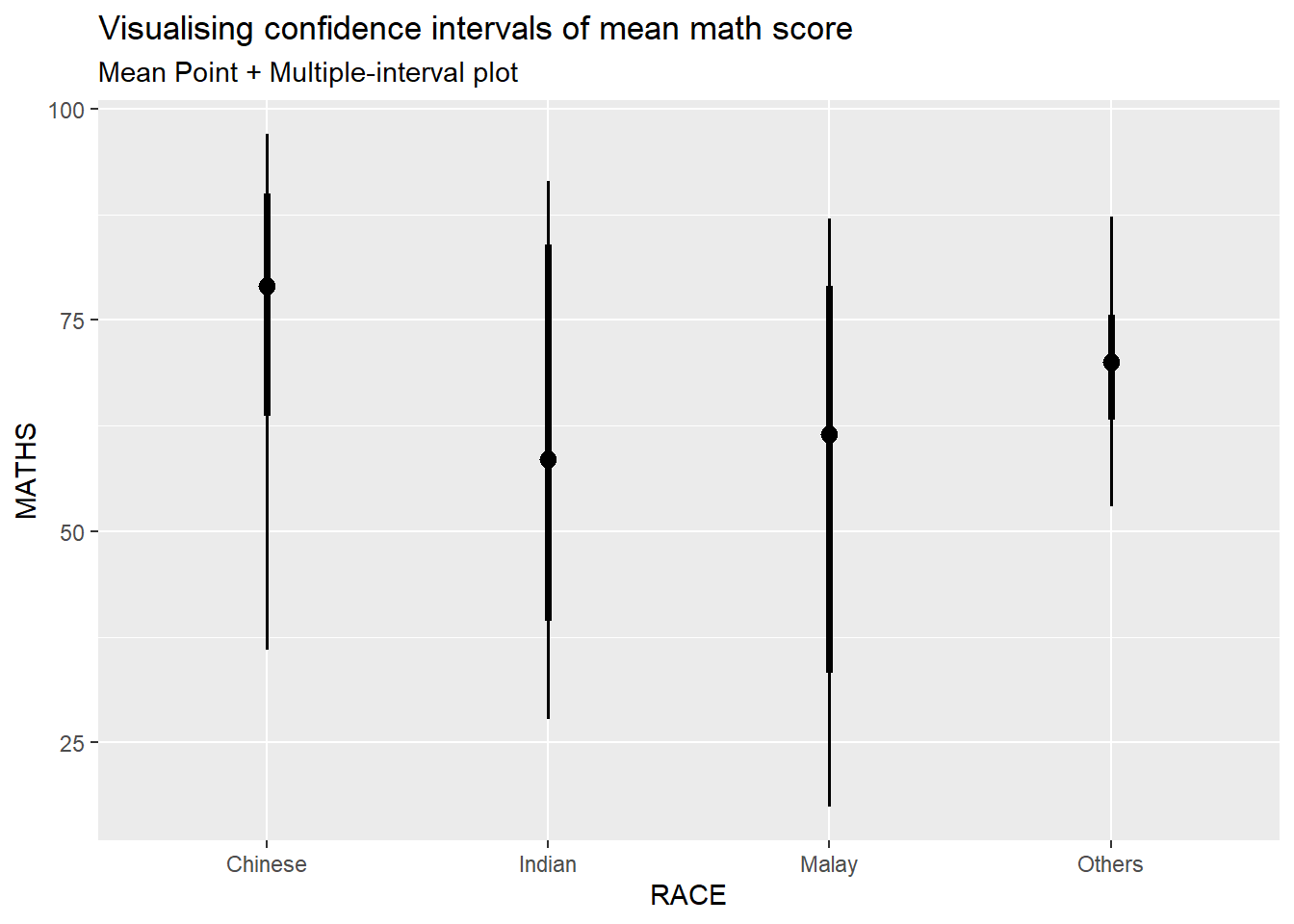
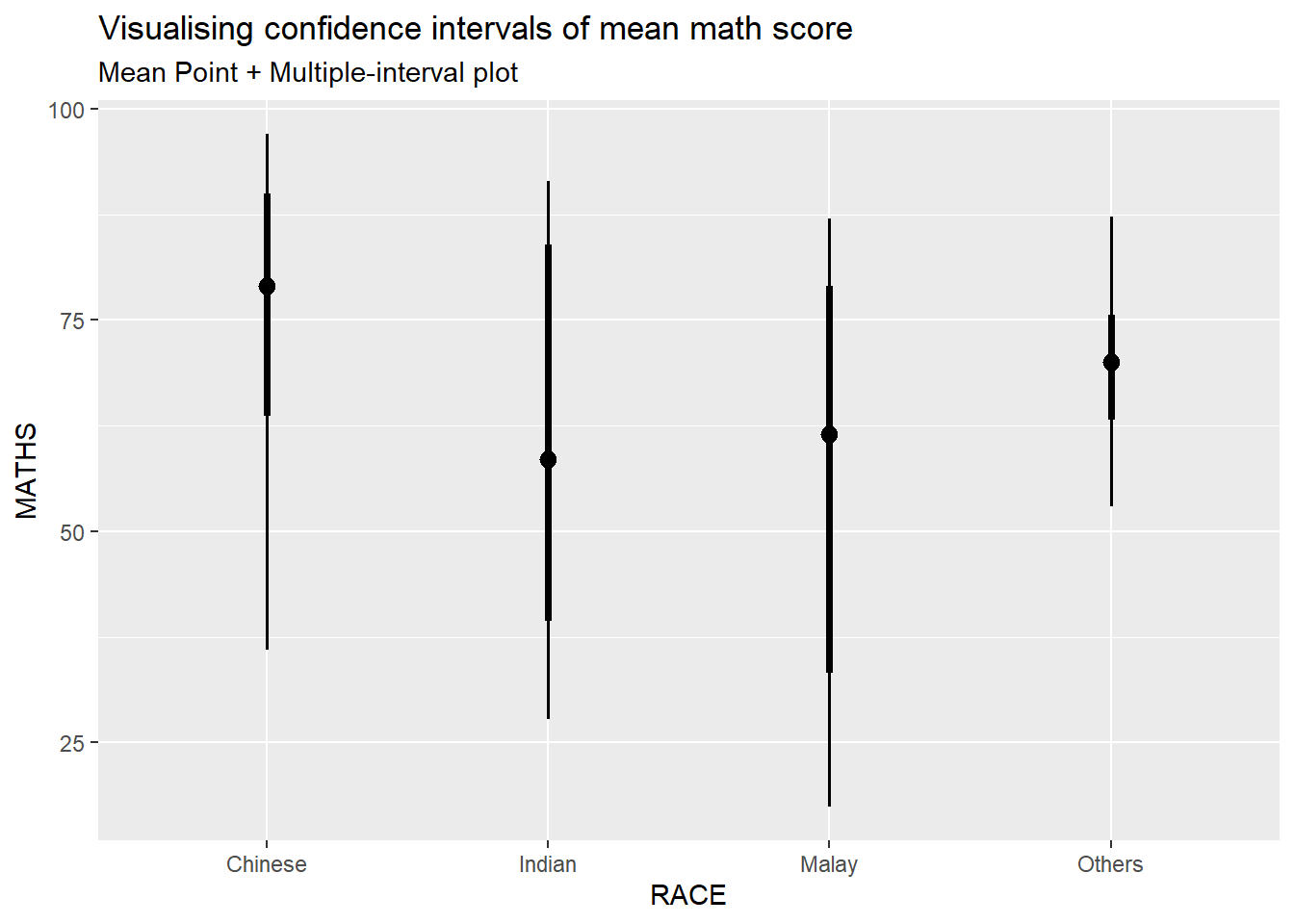
Showing 95% and 99% confidence interval with mean
exam %>%
ggplot(aes(x = RACE,
y = MATHS)) +
#refer to point_interval argument in stat_pointinterval() help
stat_pointinterval(
.point = mean,
.interval = c(qi(0.05), qi(0.01))
) +
labs(
title = "Visualising confidence intervals of mean math score",
subtitle = "Mean Point + Multiple-interval plot"
)
stat_gradientinterval() of ggdist is used to build a visual for displaying distribution of maths scores by race.
exam %>%
ggplot(aes(x = RACE,
y = MATHS)) +
stat_gradientinterval(
fill = "skyblue",
show.legend = TRUE
) +
labs(
title = "Visualising confidence intervals of mean math score",
subtitle = "Gradient + interval plot")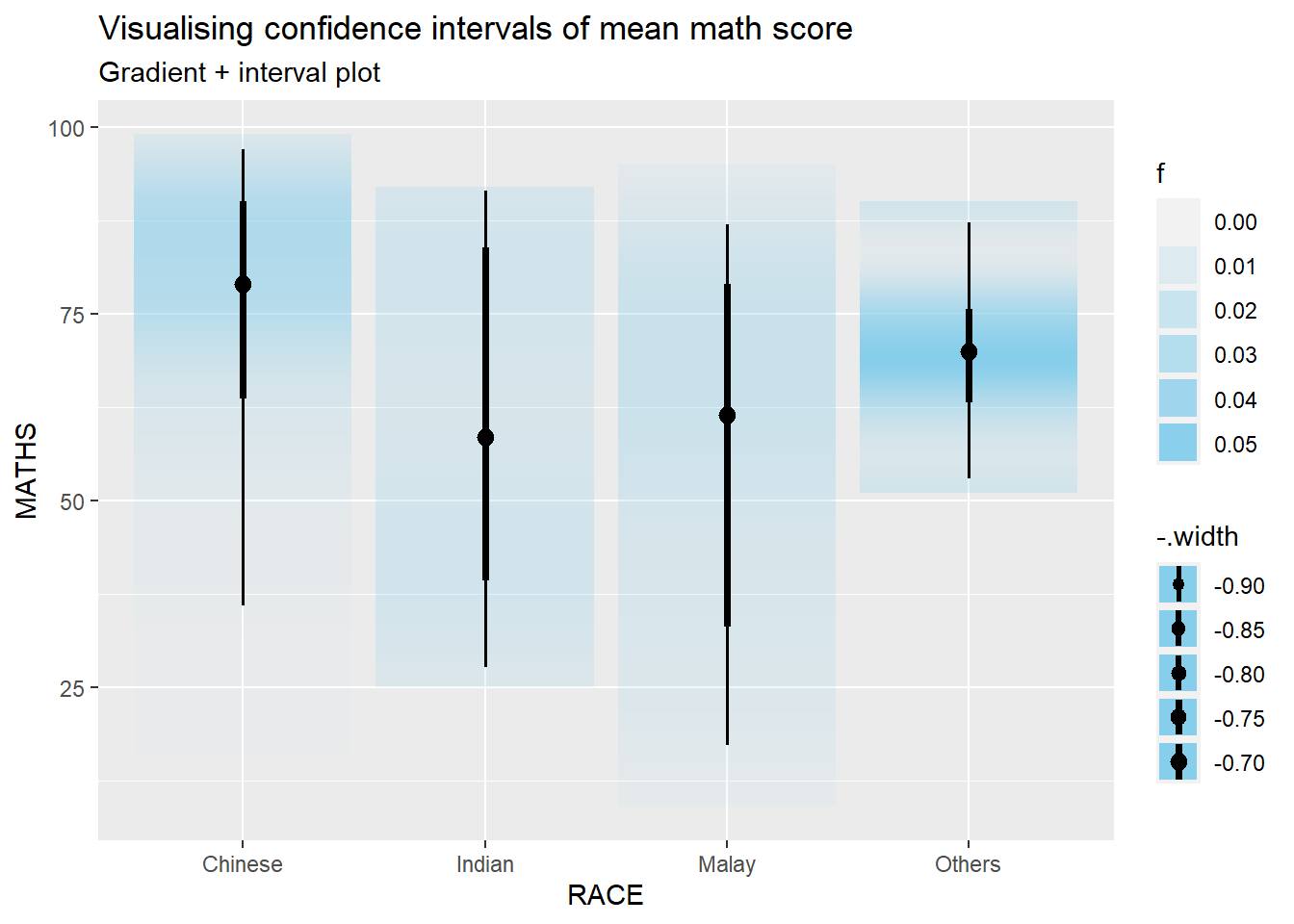
Visualising Uncertainty with Hypothetical Outcome Plots (HOPs)
#devtools::install_github("wilkelab/ungeviz", force = TRUE)
library(ungeviz)ggplot(data = exam,
(aes(x = factor(RACE), y = MATHS))) +
geom_point(position = position_jitter(
height = 0.3, width = 0.05),
size = 0.4, color = "#0072B2", alpha = 1/2) +
geom_hpline(data = sampler(25, group = RACE), height = 0.6, color = "#D55E00") +
theme_bw() +
# `.draw` is a generated column indicating the sample draw
transition_states(.draw, 1, 3)+
xlab("Race")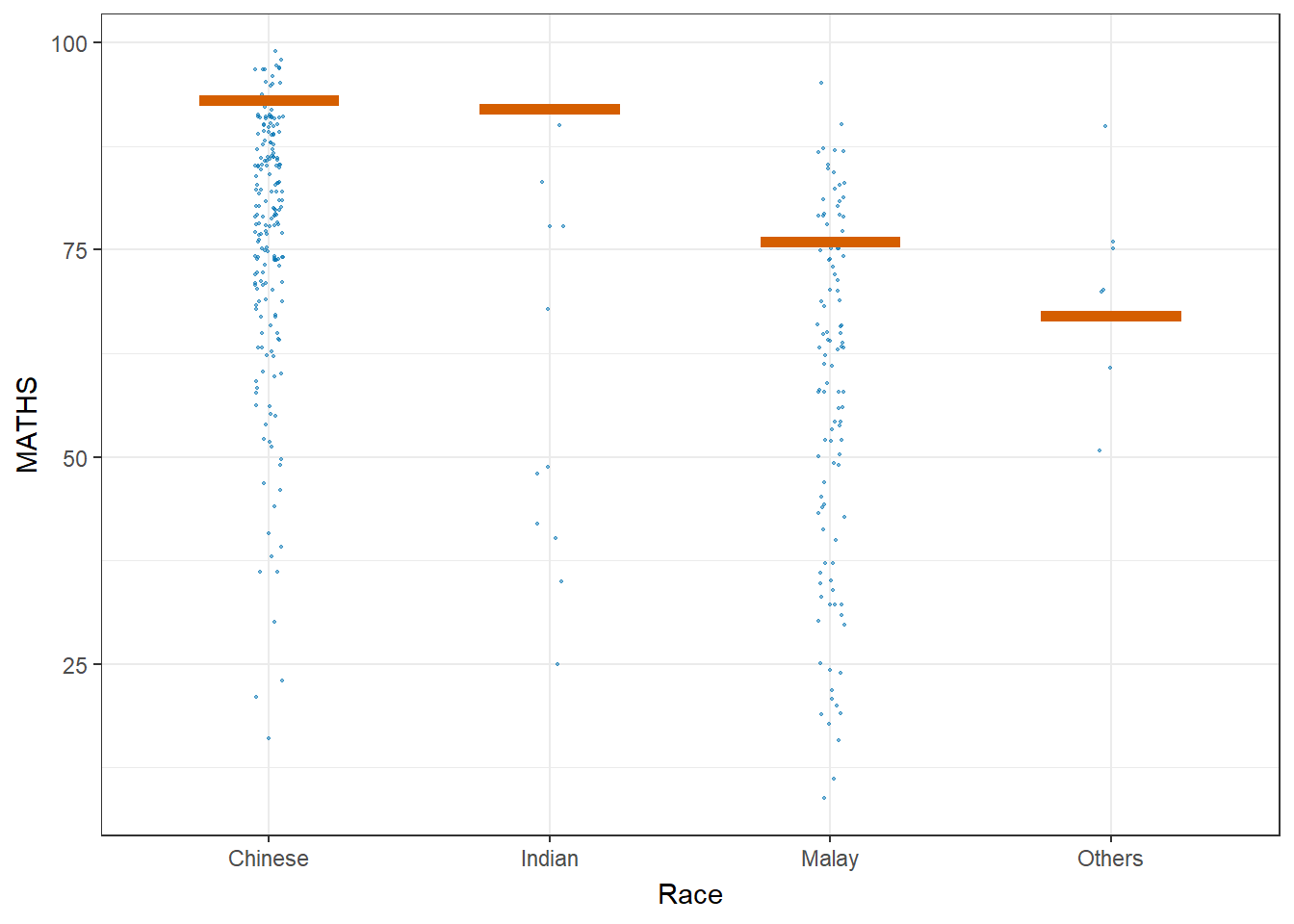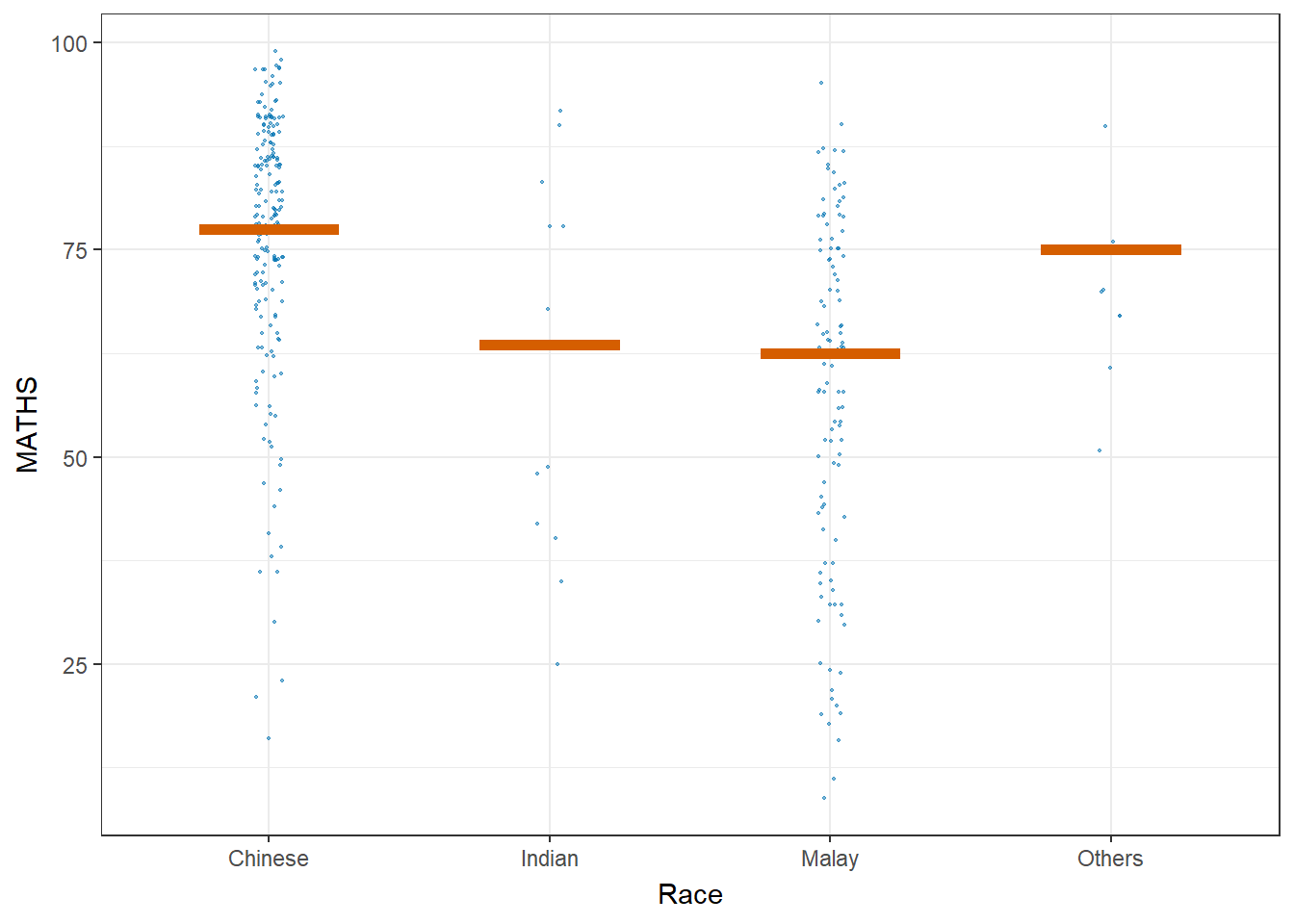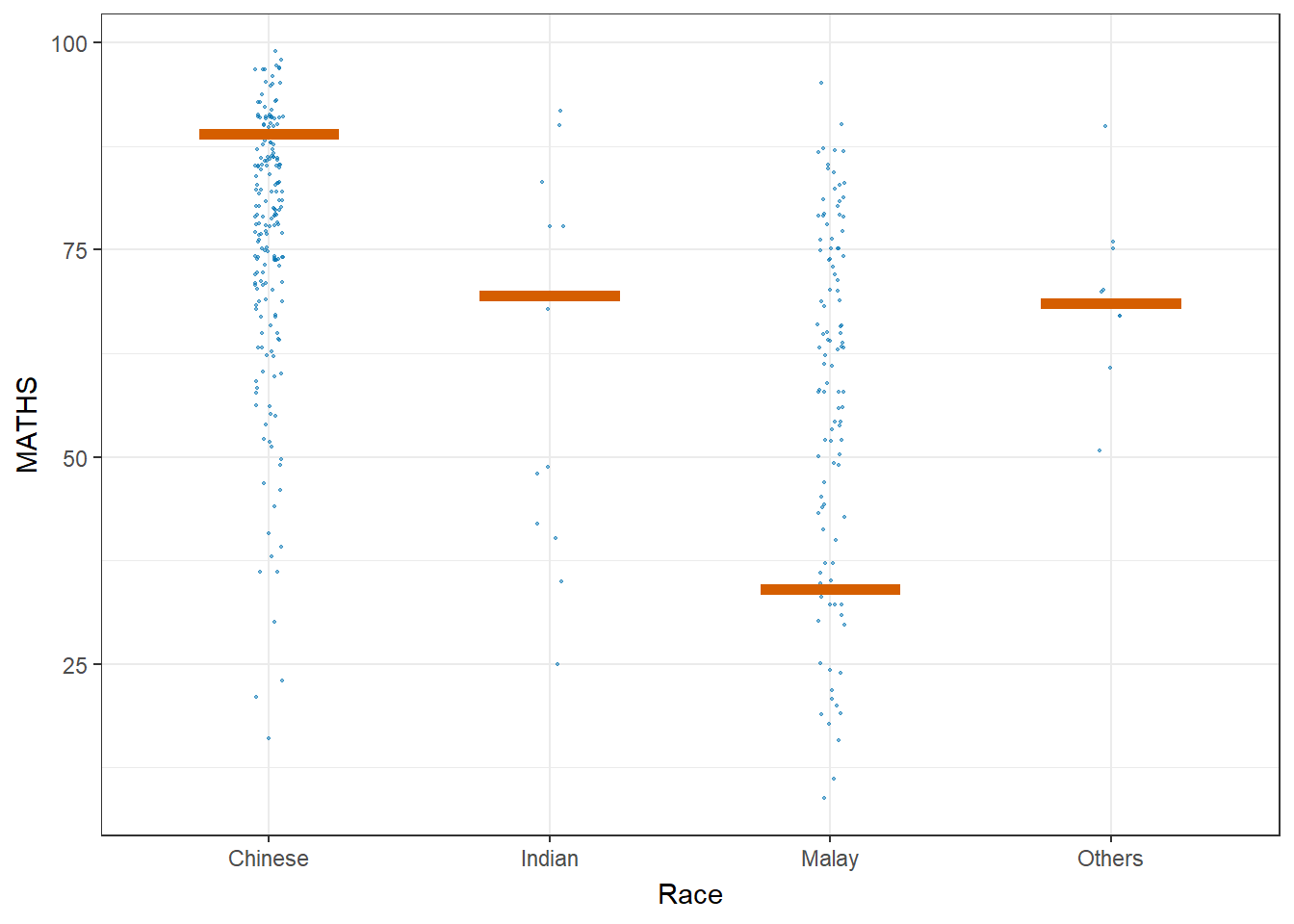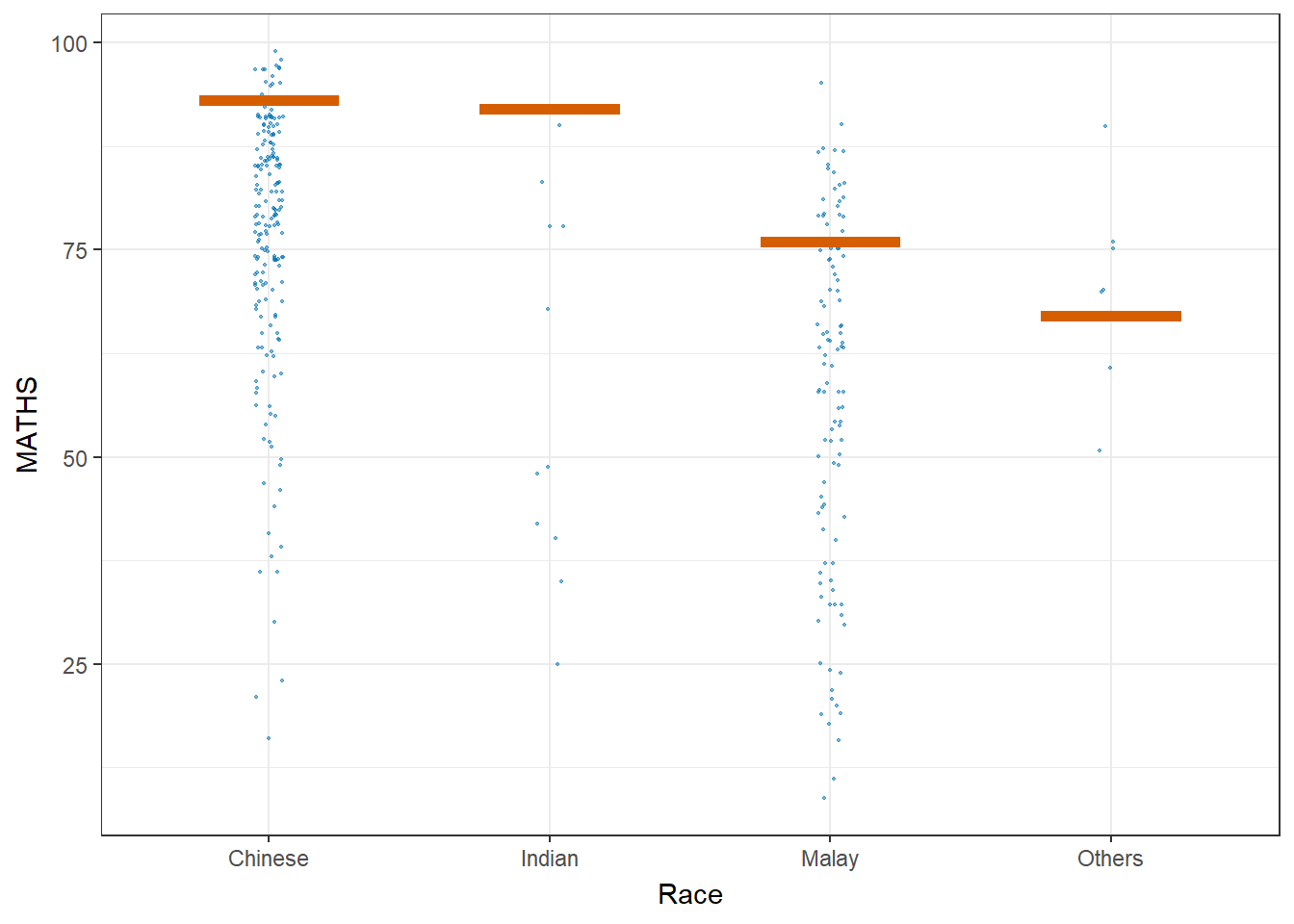
Funnel Plots for Fair Comparisons
Getting started
pacman::p_load(tidyverse, FunnelPlotR, plotly, knitr)covid19 <- read_csv("data/COVID-19_DKI_Jakarta.csv") %>%
mutate_if(is.character, as.factor)
covid19# A tibble: 267 × 7
`Sub-district ID` City District `Sub-district` Positive Recovered Death
<dbl> <fct> <fct> <fct> <dbl> <dbl> <dbl>
1 3172051003 JAKARTA U… PADEMAN… ANCOL 1776 1691 26
2 3173041007 JAKARTA B… TAMBORA ANGKE 1783 1720 29
3 3175041005 JAKARTA T… KRAMAT … BALE KAMBANG 2049 1964 31
4 3175031003 JAKARTA T… JATINEG… BALI MESTER 827 797 13
5 3175101006 JAKARTA T… CIPAYUNG BAMBU APUS 2866 2792 27
6 3174031002 JAKARTA S… MAMPANG… BANGKA 1828 1757 26
7 3175051002 JAKARTA T… PASAR R… BARU 2541 2433 37
8 3175041004 JAKARTA T… KRAMAT … BATU AMPAR 3608 3445 68
9 3171071002 JAKARTA P… TANAH A… BENDUNGAN HIL… 2012 1937 38
10 3175031002 JAKARTA T… JATINEG… BIDARA CINA 2900 2773 52
# ℹ 257 more rowsFunnelPlotR methods
FunnelPlotR package requires a numerator (events of interest), denominator (population to be considered) and group:
limit: plot limits (95 or 99). label_outliers: label outliers (true or false). Poisson_limits: add Poisson limits to the plot. OD_adjust: add overdispersed limits to the plot. xrange and yrange: specify the range to display for axes
funnel_plot(
numerator = covid19$Death,
denominator = covid19$Positive,
group = covid19$`Sub-district`,
data_type = "PR",
xrange = c(0, 6500),
yrange = c(0, 0.05),
label = NA,
title = "Cumulative COVID-19 Fatality Rate by Cumulative Total Number of COVID-19 Positive Cases", #<<
x_label = "Cumulative COVID-19 Positive Cases", #<<
y_label = "Cumulative Fatality Rate" #<<
)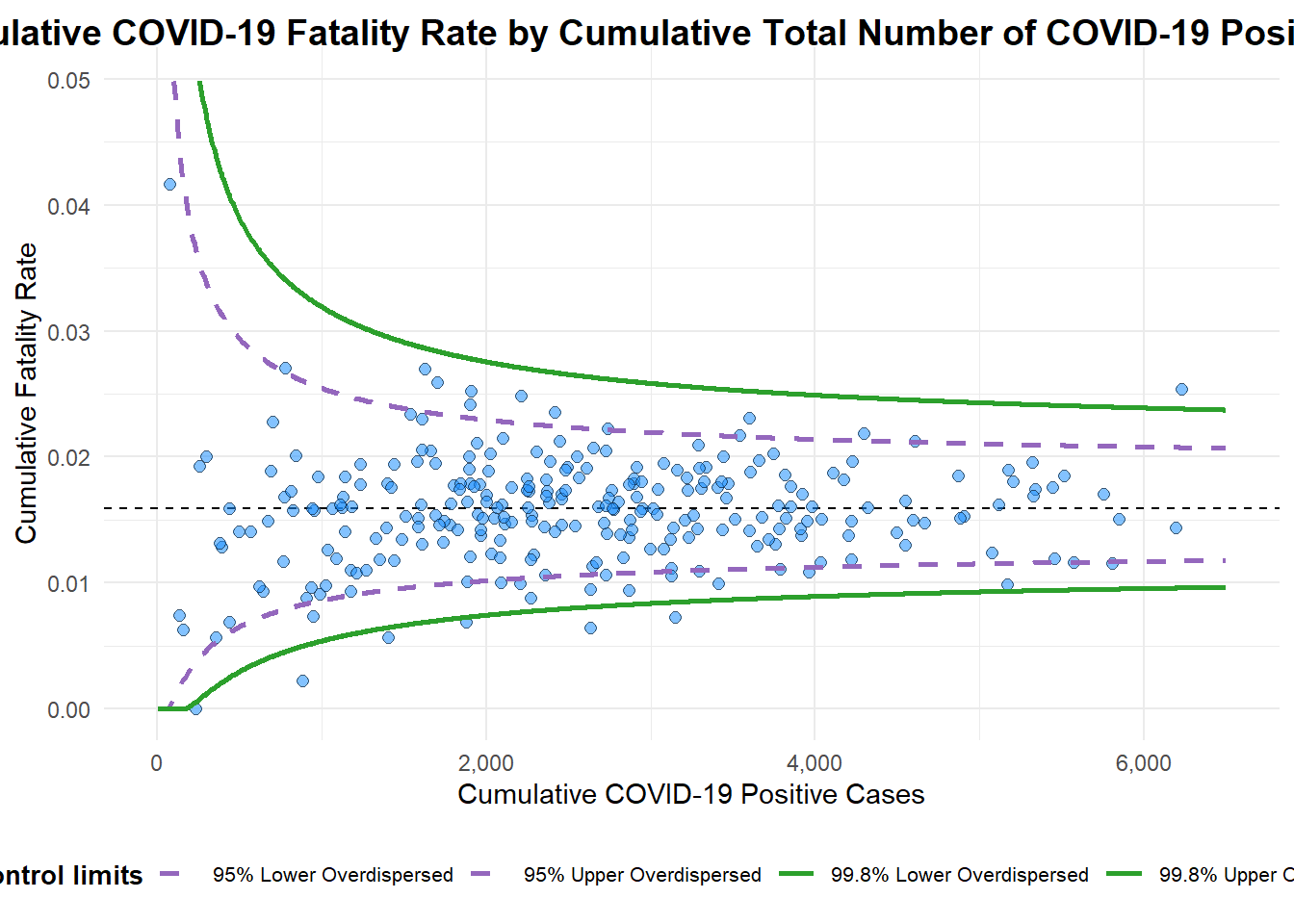
A funnel plot object with 267 points of which 7 are outliers.
Plot is adjusted for overdispersion. Funnel Plot for Fair Visual Comparison: ggplot2 methods
Derive cumulative death rate and standard error of cumulative death rate.
df <- covid19 %>%
mutate(rate = Death / Positive) %>%
mutate(rate.se = sqrt((rate*(1-rate)) / (Positive))) %>%
filter(rate > 0)Compute fit.mean
fit.mean <- weighted.mean(df$rate, 1/df$rate.se^2)Calculate lower and upper limits for 95% and 99.9% CI
number.seq <- seq(1, max(df$Positive), 1)
number.ll95 <- fit.mean - 1.96 * sqrt((fit.mean*(1-fit.mean)) / (number.seq))
number.ul95 <- fit.mean + 1.96 * sqrt((fit.mean*(1-fit.mean)) / (number.seq))
number.ll999 <- fit.mean - 3.29 * sqrt((fit.mean*(1-fit.mean)) / (number.seq))
number.ul999 <- fit.mean + 3.29 * sqrt((fit.mean*(1-fit.mean)) / (number.seq))
dfCI <- data.frame(number.ll95, number.ul95, number.ll999,
number.ul999, number.seq, fit.mean)Plot a static funnel plot
p <- ggplot(df, aes(x = Positive, y = rate)) +
geom_point(aes(label=`Sub-district`),
alpha=0.4) +
geom_line(data = dfCI,
aes(x = number.seq,
y = number.ll95),
size = 0.4,
colour = "grey40",
linetype = "dashed") +
geom_line(data = dfCI,
aes(x = number.seq,
y = number.ul95),
size = 0.4,
colour = "grey40",
linetype = "dashed") +
geom_line(data = dfCI,
aes(x = number.seq,
y = number.ll999),
size = 0.4,
colour = "grey40") +
geom_line(data = dfCI,
aes(x = number.seq,
y = number.ul999),
size = 0.4,
colour = "grey40") +
geom_hline(data = dfCI,
aes(yintercept = fit.mean),
size = 0.4,
colour = "grey40") +
coord_cartesian(ylim=c(0,0.05)) +
annotate("text", x = 1, y = -0.13, label = "95%", size = 3, colour = "grey40") +
annotate("text", x = 4.5, y = -0.18, label = "99%", size = 3, colour = "grey40") +
ggtitle("Cumulative Fatality Rate by Cumulative Number of COVID-19 Cases") +
xlab("Cumulative Number of COVID-19 Cases") +
ylab("Cumulative Fatality Rate") +
theme_light() +
theme(plot.title = element_text(size=12),
legend.position = c(0.91,0.85),
legend.title = element_text(size=7),
legend.text = element_text(size=7),
legend.background = element_rect(colour = "grey60", linetype = "dotted"),
legend.key.height = unit(0.3, "cm"))
p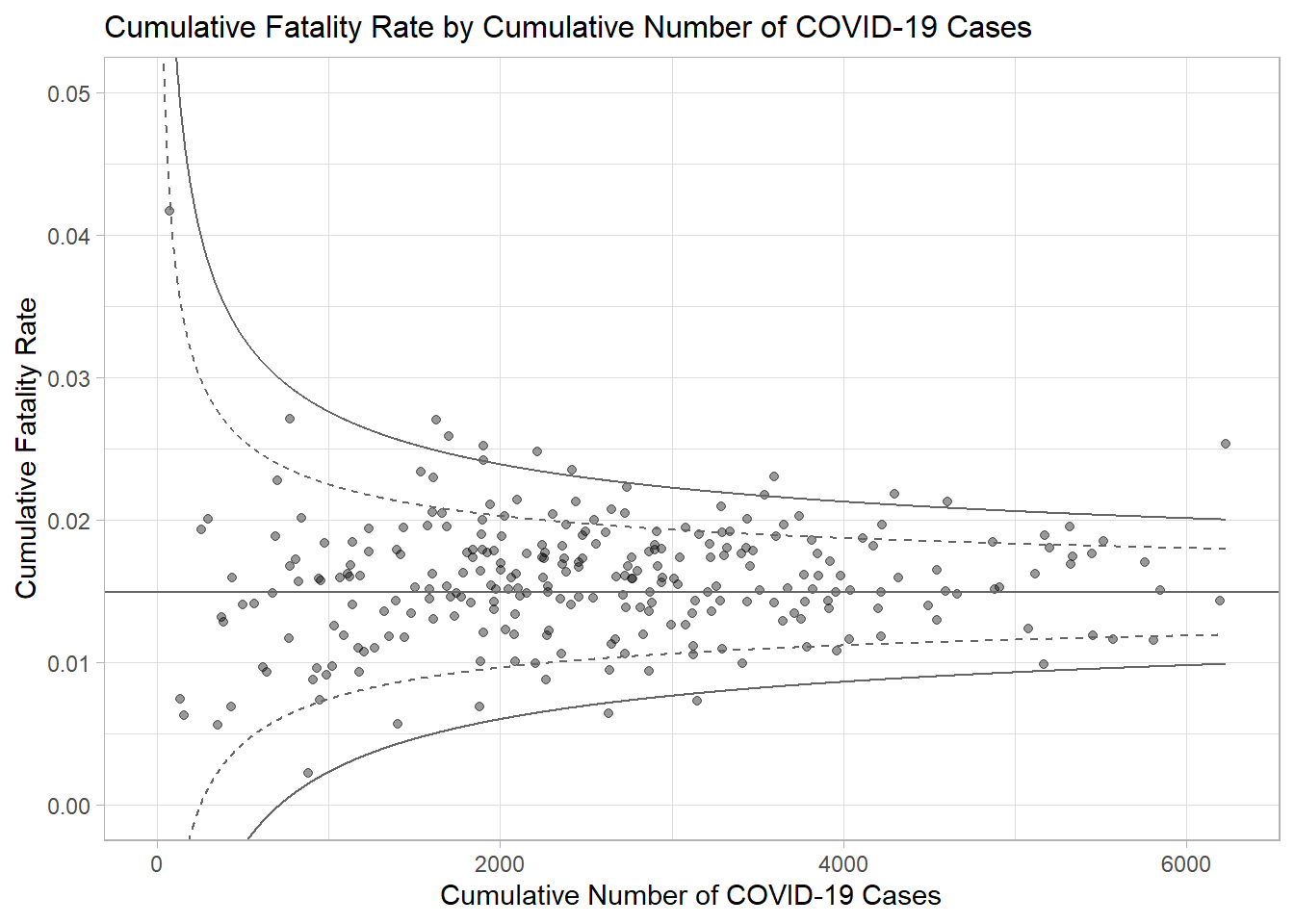
Plot Interactive funnel plot
Use ggplotly() of plotly r package.
fp_ggplotly <- ggplotly(p,
tooltip = c("label",
"x",
"y"))
fp_ggplotly